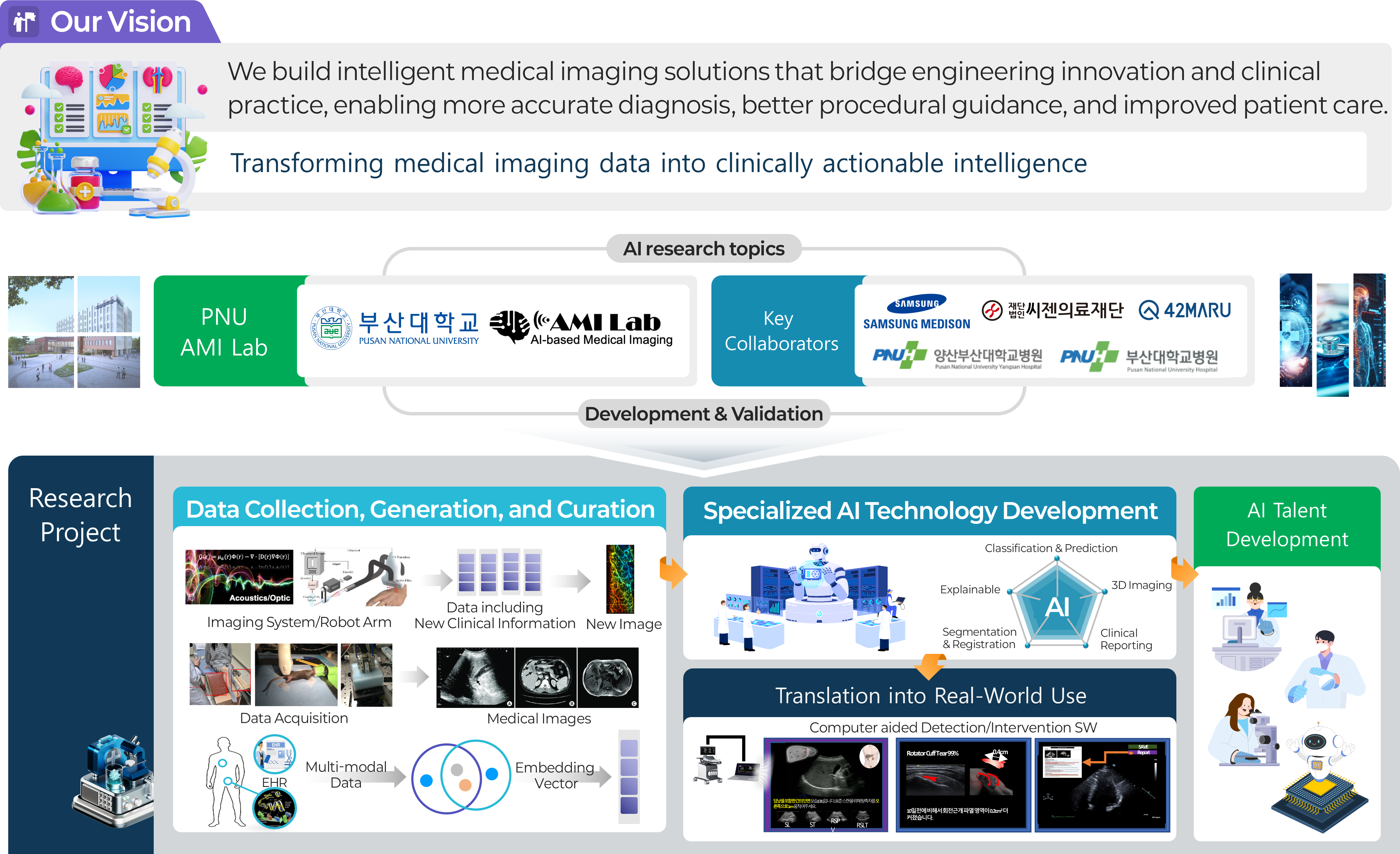
Pusan National University AMILab (AI Medical Imaging Lab) is a research laboratory dedicated to next-generation medical artificial intelligence based on medical devices and healthcare data. In particular, we develop diagnostic, analytical, and guidance technologies that can support real clinical practice by leveraging a wide range of medical imaging modalities and biosignals, including ultrasound, photoacoustic imaging, CT, and MRI. AMILab goes beyond simply improving the performance of deep learning models. We pursue AI methodologies that take into account both the physical principles of medical devices and the practical needs of clinical care. Through this approach, our goal is not limited to laboratory-level algorithm development, but extends to building clinically meaningful intelligent medical imaging technologies that can be truly used by healthcare professionals. Our major research areas include ultrasound and photoacoustic image reconstruction, 3D/4D medical image analysis, disease prediction based on biosignals and multimodal health screening data, real-time guidance systems for assisting clinicians, robotic-arm-based automated ultrasound scanning and diagnosis, and vision-language-model (VLM)-based medical AI that integrates medical images with radiology reports and clinical text. These research topics span medical image processing, signal processing, deep learning, computer vision, and multimodal learning, while pursuing both engineering rigor and clinical applicability. In addition, AMILab conducts translational research aimed at solving real problems in healthcare through close collaboration with clinicians, domestic and international research institutions, and industry partners. In collaboration with Samsung Medison, Seegene Medical Foundation, 42Maru, Pusan National University Hospital, and Pusan National University Yangsan Hospital, we are building a research ecosystem that connects data acquisition, algorithm development, clinical validation, and real-world deployment. Through these efforts, we aim to produce top-tier research outcomes in SCI-indexed journals and leading venues such as MICCAI and IEEE conferences, while also advancing our technologies toward actual medical devices and clinical environments. Research at AMILab follows a full translational pipeline. We first establish reliable foundations through the collection, generation, and curation of medical data for AI training. Building on this foundation, we develop and validate specialized AI technologies tailored to specific medical problems. Ultimately, we seek to expand these technologies into real clinical practice through clinical validation and real-world deployment. Through this entire process, AMILab is committed not only to developing advanced technologies, but also to training AI talent capable of identifying important problems, solving them with engineering approaches, and connecting them to genuine clinical value. AMILab provides an excellent research environment for students who aspire to define meaningful problems on their own, solve them through engineering, and ultimately contribute to healthcare. We welcome students interested in medical AI, medical imaging, computer vision, deep learning, signal processing, multimodal AI, VLMs, and medical robotics.
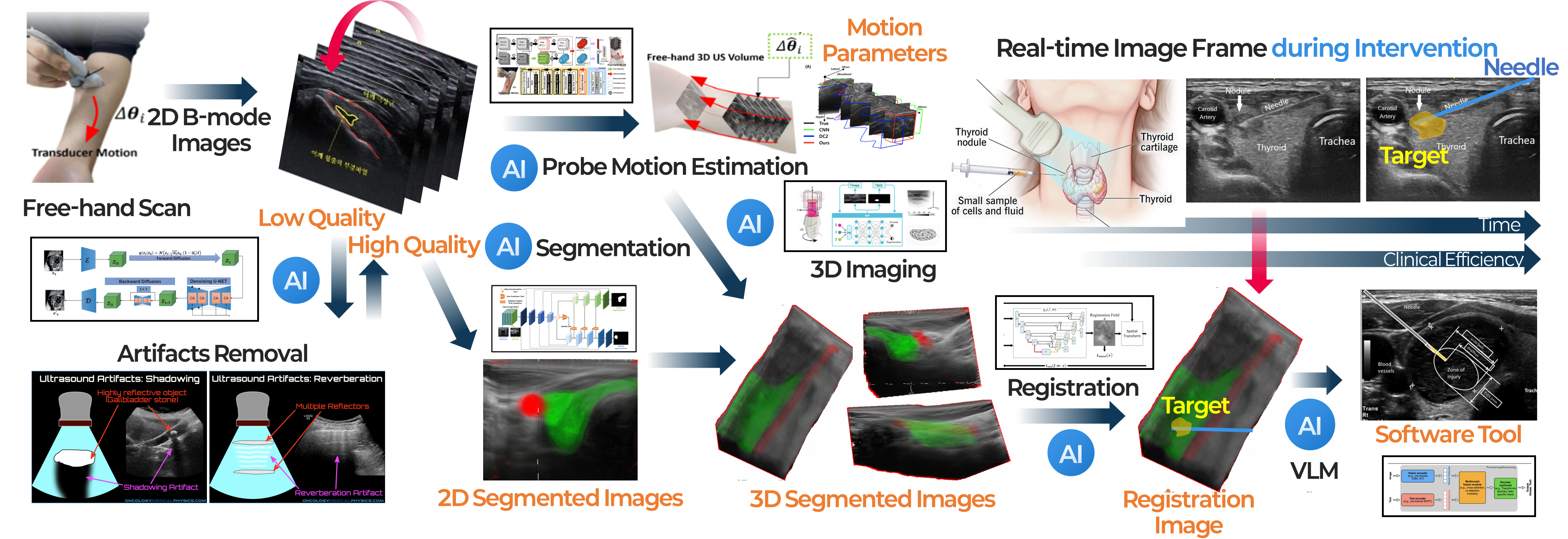
This project focuses on developing AI technologies to reconstruct 3D ultrasound images from 2D ultrasound scans acquired in a free-hand manner, without relying on external position sensors. Conventional free-hand 3D ultrasound systems typically require additional tracking devices to estimate the probe’s position and orientation, but such systems can increase complexity and cost while reducing flexibility in real clinical settings. To overcome these limitations, we are developing a deep learning-based framework that estimates probe motion directly from consecutively acquired 2D ultrasound images and reconstructs a 3D ultrasound volume from them. By incorporating spatial continuity during scanning and anatomical consistency of the target structure, our method aims to achieve more accurate and stable 3D reconstruction. This approach enables practical and user-friendly 3D ultrasound imaging without external sensors, and can be further extended by integrating image segmentation techniques to selectively render organs, vessels, and lesion regions in 3D. Ultimately, we aim to develop this technology into an assistive software tool that can support disease assessment, procedural and surgical planning, and intra-procedural image guidance. This project represents one of the core ultrasound AI research topics at AMILab, Pusan National University, and reflects our broader goal of combining medical imaging physics with artificial intelligence to create next-generation imaging technologies for real clinical use. Related studies have led to research outcomes in international journals and top-tier conferences, and the project is currently supported by the Basic Science Research Program of the National Research Foundation of Korea (NRF) and the AI Convergence Innovation Human Resources Development Program of the Institute of Information & Communications Technology Planning & Evaluation (IITP). In addition, accurate probe motion estimation and high-quality 3D reconstruction strongly depend on the quality of the input 2D ultrasound images. For this reason, we are also pursuing a closely related research direction under the support of Samsung Medison to generate high-quality 2D ultrasound images even with limited ultrasound transmissions and receptions.
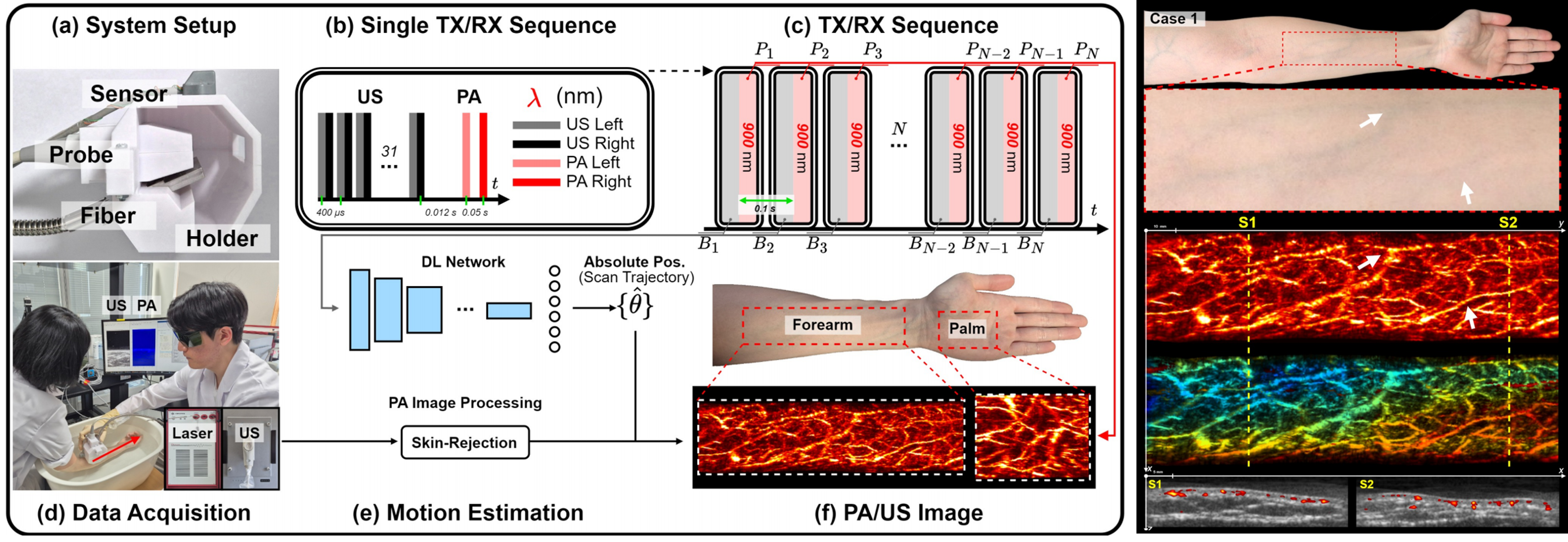
This project focuses on developing AI and imaging technologies that combine conventional ultrasound systems with laser devices to acquire photoacoustic images and extend them toward practical clinical use. Photoacoustic imaging is a promising modality that combines the advantages of light and ultrasound to visualize functional and vascular information in biological tissues. In particular, it enables noninvasive observation of microvascular structures and blood-related information. However, conventional photoacoustic imaging systems often involve complex hardware configurations, have limitations in imaging range and penetration depth, and face challenges in obtaining stable 3D images in free-hand scanning settings, which has hindered their broader clinical adoption. To overcome these limitations, we developed a free-hand 3D photoacoustic imaging technique that tracks probe position and motion using ultrasound images and reconstructs 3D microvascular structures by registering and accumulating simultaneously acquired photoacoustic images. In particular, by incorporating our free-hand 3D technology that estimates probe motion from ultrasound images without external position sensors, we aimed to enable a more practical and user-friendly approach to 3D photoacoustic imaging. This approach makes it possible to move beyond 2D cross-sectional photoacoustic vascular images and reconstruct them in 3D, providing a more intuitive and volumetric representation of microvascular structures. Furthermore, this research goes beyond simple 3D reconstruction and is evolving toward broader clinical applications of photoacoustic imaging. In particular, we are also exploring technical approaches to improve imaging depth so that useful information can be obtained from deeper tissues, thereby extending the technology beyond superficial vascular imaging toward a wider range of clinical applications. This project is currently supported by the Basic Science Research Program of the National Research Foundation of Korea (NRF). In the long term, we aim to advance this work into a next-generation photoacoustic imaging platform that can support microvascular imaging, functional imaging, and applications in diagnosis and therapy guidance.
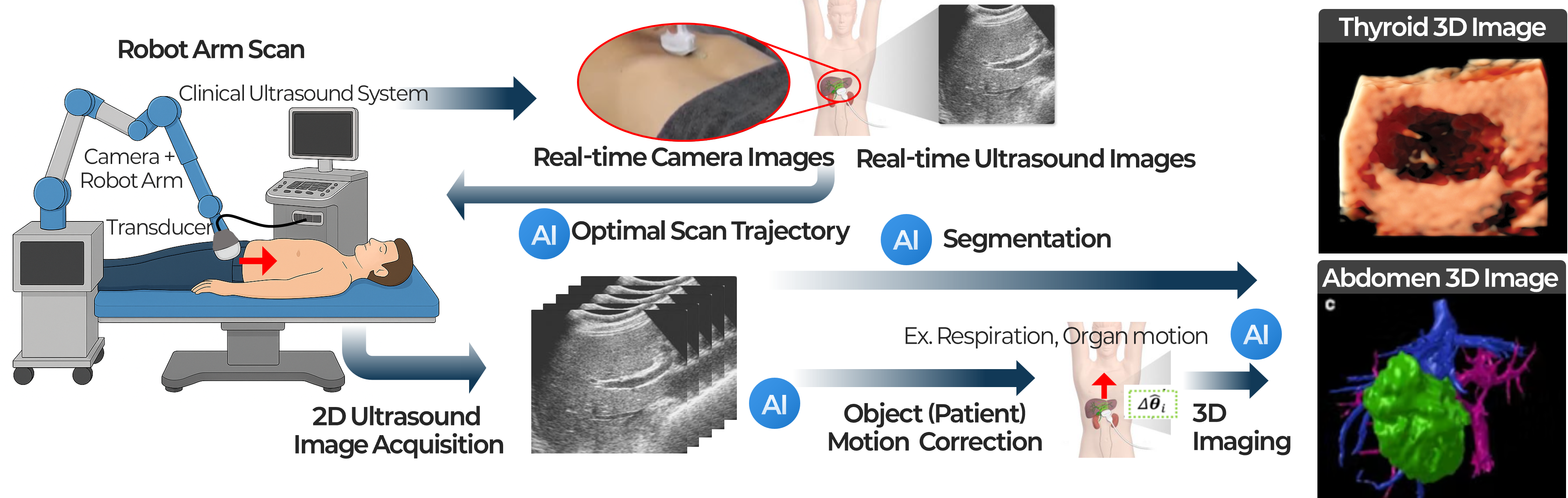
This project focuses on developing AI and robotics technologies for more automated and consistent ultrasound scanning using a robotic arm and cameras. Conventional ultrasound examinations rely heavily on the operator’s skill and manual technique, which means that probe position, angle, applied pressure, and scanning path can vary even when examining the same patient. Such variability affects image quality and reproducibility, and can become a significant limitation in clinical scenarios where longitudinal follow-up or quantitative comparison is important. Although this approach requires a higher system cost, our goal is to reduce the inconsistency of manual scanning and enable more standardized ultrasound acquisition through automated scanning technology that combines a robotic arm and cameras. To achieve this, we are developing camera-based computer vision techniques to recognize the patient’s pose and anatomical location and to automatically identify appropriate scanning regions. In addition, the robotic arm is designed to adjust the probe position and orientation appropriately by jointly using patient responses and movements observed through cameras, together with real-time ultrasound images acquired from the ultrasound imaging system. In this way, our system goes beyond simply repeating pre-defined trajectories and aims to enable intelligent ultrasound scanning that can flexibly respond to patient condition and image quality. In particular, because the robotic arm can precisely control and record the probe’s position and orientation, it provides a strong foundation for acquiring high-quality 3D ultrasound images. At the same time, deep learning models can focus less on estimating the probe position itself and more on compensating for errors caused by subtle patient motion, respiration, and soft-tissue deformation, enabling more precise and stable 3D reconstruction and automated diagnostic assistance. Ultimately, we aim to advance this technology into an integrated intelligent ultrasound platform that encompasses automated ultrasound scanning, standard-plane acquisition, 3D imaging, and support for diagnosis and interventions. This project is supported by the Basic Science Research Program of the National Research Foundation of Korea (NRF) and the AI Convergence Innovation Human Resources Development Program of the Institute of Information & Communications Technology Planning & Evaluation (IITP).
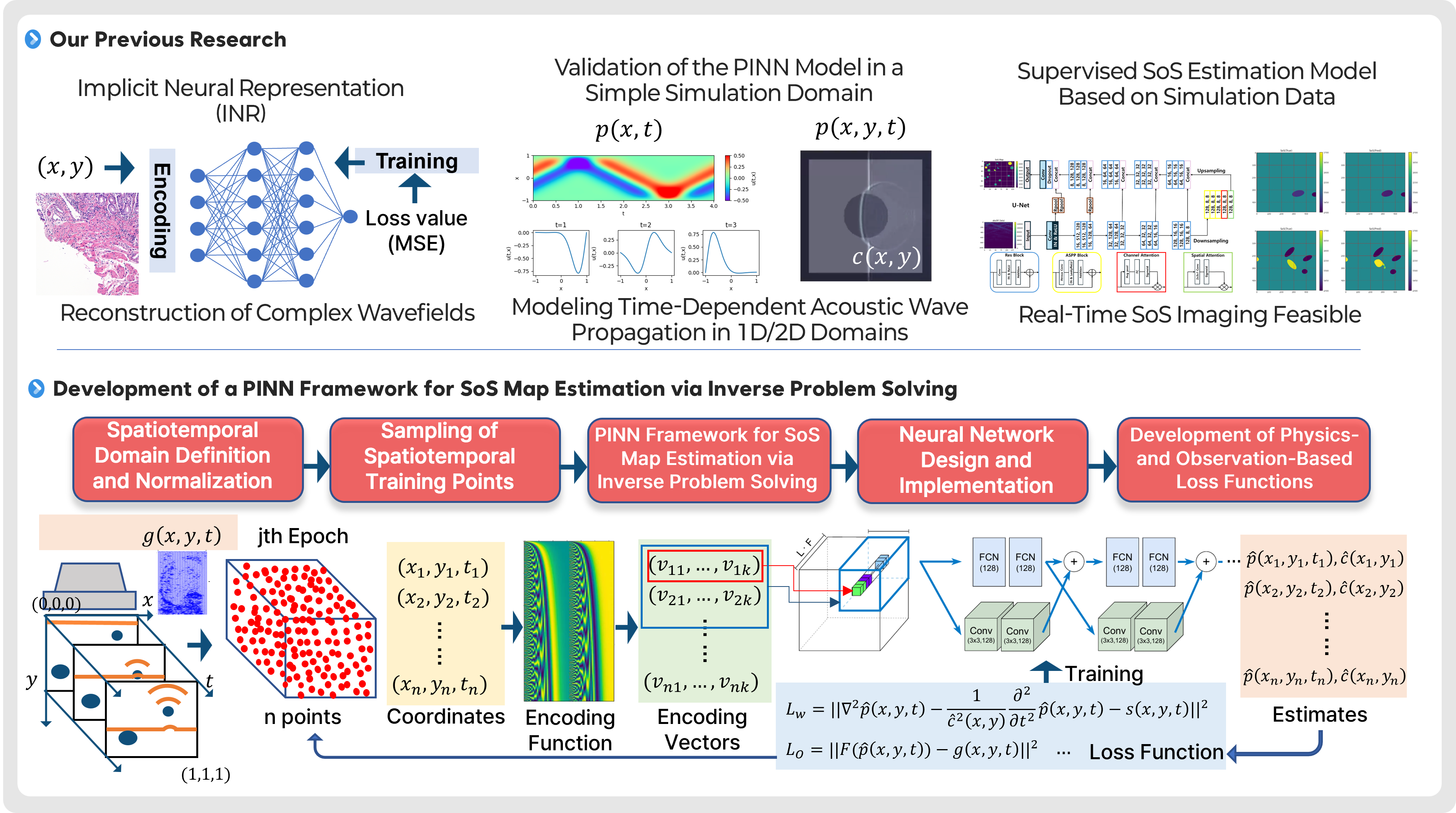
This project focuses on developing AI technologies to estimate and image the internal speed-of-sound (SoS) distribution in tissue using physics-informed neural networks (PINNs). In ultrasound imaging, the speed of sound is a key physical parameter that governs wave propagation and can vary depending on tissue type and condition. Therefore, SoS maps can provide additional tissue property information that is difficult to obtain from conventional B-mode images alone, and may serve as a new diagnostic indicator in the future. However, estimating the SoS distribution is a representative inverse problem, in which internal material properties must be inferred from observed ultrasound signals. This problem is computationally complex and highly sensitive to noise and variations in measurement conditions, making stable reconstruction challenging. To address this challenge, we are developing a PINN-based framework that simultaneously incorporates the physical laws governing acoustic wave propagation and actual observed data. We first reproduce time-dependent acoustic wave propagation in relatively simple 1D and 2D simulation domains to validate the PINN model and its training strategies. Building on this foundation, we aim to establish a full learning framework for SoS map estimation through spatiotemporal domain definition and normalization, sampling of training points, development of differentiable coordinate encoding methods, neural network design, and loss functions that jointly reflect physical residuals and observation errors. Through this approach, we seek to move beyond conventional supervised estimation and achieve more physically consistent and generalizable SoS reconstruction. Ultimately, we aim to advance this technology into a new ultrasound-based quantitative imaging modality that can estimate SoS images in real time or near real time and be extended toward clinical use. In the long term, we plan to incorporate more complex tissue environments, real ultrasound acquisition conditions, and diverse lesion scenarios, with the goal of developing a next-generation ultrasound imaging platform that provides functional and material-property information complementary to conventional structural imaging. This project is supported by the Basic Science Research Program of the National Research Foundation of Korea (NRF) and the AI Convergence Innovation Human Resources Development Program of the Institute of Information & Communications Technology Planning & Evaluation (IITP).
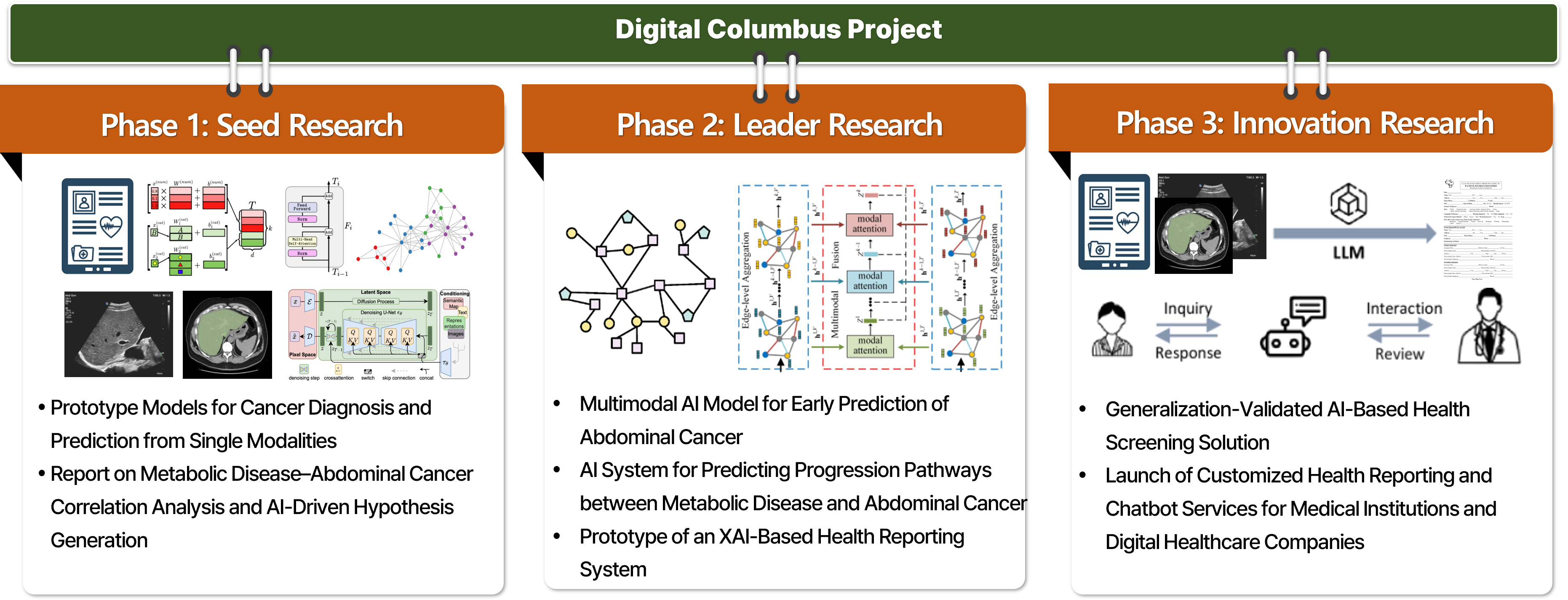
This project focuses on developing explainable AI models that analyze the correlations between metabolic diseases and abdominal cancers and predict disease progression using multimodal data accumulated through health screening programs. In real clinical settings, metabolic diseases and cancer are likely to interact with each other over long periods rather than existing independently. However, technologies that systematically analyze these relationships and leverage them for early prediction are not yet well established. Our goal is to integratively utilize imaging data, clinical measurements, questionnaire information, and longitudinal follow-up data from health screening to better understand inter-disease relationships and progression pathways, and ultimately expand this work into clinically applicable AI-based health management technologies. This project is being carried out in multiple stages. In the current Phase 1: Seed Research, we focus on extracting longitudinal features from repeated health screening data collected at Pusan National University Yangsan Hospital and analyzing the relationships and temporal changes among metabolic disease-related indicators. Based on this foundation, we aim to build prototype models for disease diagnosis and prediction at the single-modality level, while establishing the data foundation and initial hypotheses for future complex disease analysis. In Phase 2: Leader Research, we plan to explicitly connect metabolic diseases with abdominal cancers and expand the work into multimodal AI models for early abdominal cancer prediction and disease progression pathway prediction. In Phase 3: Innovation Research, we aim to advance the project into service-level AI health screening solutions, including explainable AI (XAI)-based health reporting systems applicable to real medical institutions and digital healthcare companies. This research is currently being conducted through collaboration with the Department of Family Medicine at Pusan National University Yangsan Hospital and multiple research teams at the AI Convergence Research Center of Pusan National University. It represents a convergent translational research effort that jointly considers medical data interpretation, AI modeling, and clinical applicability. This project is supported by the Digital Columbus Project of the Institute of Information & Communications Technology Planning & Evaluation (IITP). In the long term, we aim to develop this work into a next-generation explainable AI platform that can enable early prediction of disease risks from health screening data and expand into personalized health management and digital healthcare services.
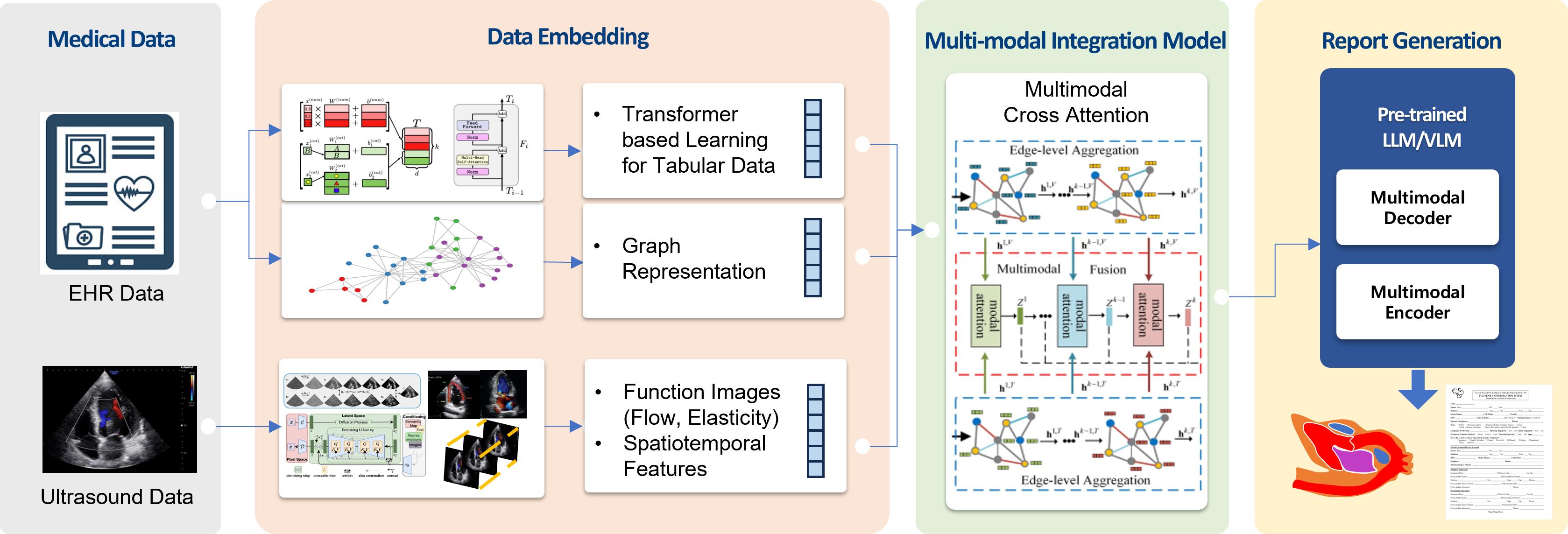
This project focuses on developing an AI platform that automatically generates echocardiography reports by extracting measurements from diverse echocardiographic images and integrating both imaging and numerical information. Echocardiography is an essential examination for evaluating cardiac structure and function. However, in real clinical practice, reviewing multiple imaging modes and standard views, obtaining the required measurements from each view, and synthesizing them into a final diagnostic report require substantial time and effort. This process can reduce the efficiency of examination and interpretation, and may also lead to variability in reporting consistency depending on the experience of the operator and reader. To address this challenge, we are developing an AI-based platform that takes echocardiographic images and measured values as input and automatically generates diagnostic reports. We are currently collecting tens of thousands of echocardiography cases, and based on image data and structured measurements, we are building models that combine vision-language models (VLMs) with medical knowledge to generate clinically natural and useful reports. Furthermore, we aim to extend this work toward models that can automatically generate reports directly from echocardiographic images alone, without relying on explicit numerical measurements, thereby enabling a higher level of automation and real-world applicability. This research is being conducted in collaboration with the Cardiology team at Pusan National University Hospital and represents a strong example of translational research connecting medical image interpretation, generative AI, and automated clinical reporting. Ultimately, we aim to advance this work into an intelligent reporting platform that supports the entire echocardiography workflow, helping reduce reporting time, improve report quality, and enhance usability in real clinical settings. This project is supported by the Generative AI Core Advanced Talent Development Program of the Institute of Information & Communications Technology Planning & Evaluation (IITP).
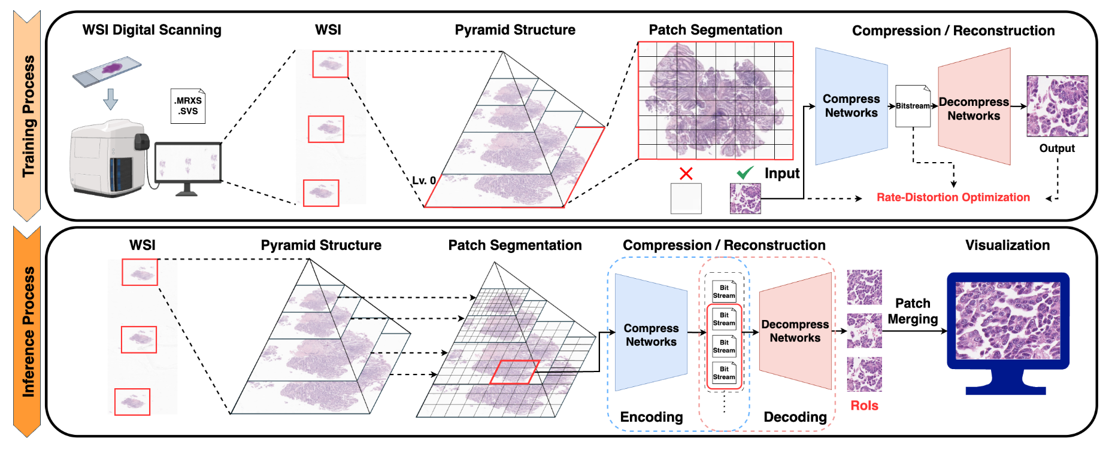
This project focuses on developing deep learning-based compression technologies for whole slide pathology images (WSIs) to enable more efficient storage, transmission, and utilization of ultra-high-resolution pathology data. In digital pathology, a single whole slide image can be extremely large, creating substantial burdens in storage, network transfer, remote diagnosis, and downstream AI analysis. Although conventional compression standards such as JPEG and JPEG2000 are widely used, they may have limitations in achieving high compression efficiency while fully preserving the fine structures and diagnostically important information contained in pathology images. Our goal is therefore to develop next-generation AI-based pathology image compression methods that reflect the unique characteristics of pathology data and maintain clinically important details more effectively. To this end, we are investigating deep learning-based encoder-decoder frameworks that transform pathology image patches into compact latent representations and reconstruct them efficiently. In parallel, we are also exploring implicit neural representation (INR)-based approaches for more continuous and expressive image modeling. While encoder-decoder architectures provide a practical and direct framework for learned compression, INR-based methods offer new possibilities for representing the fine spatial details of ultra-high-resolution images in a coordinate-based manner. Through these complementary approaches, we aim to optimize not only compression ratio and reconstruction quality, but also the preservation of structural information that is critical for pathological diagnosis. This research is being conducted in collaboration with and supported by the Seegene Medical Foundation, and represents a translational effort that considers both real pathology data and clinical applicability. Ultimately, we aim to develop this work into a practical compression solution that can be integrated into digital pathology workflows, improving storage and transmission efficiency while facilitating remote diagnosis, AI analysis, and interoperability with digital pathology platforms. In the long term, we plan to extend the technology to be compatible with various pathology image formats and viewer environments, enabling its adoption in real medical institutions and digital healthcare settings.
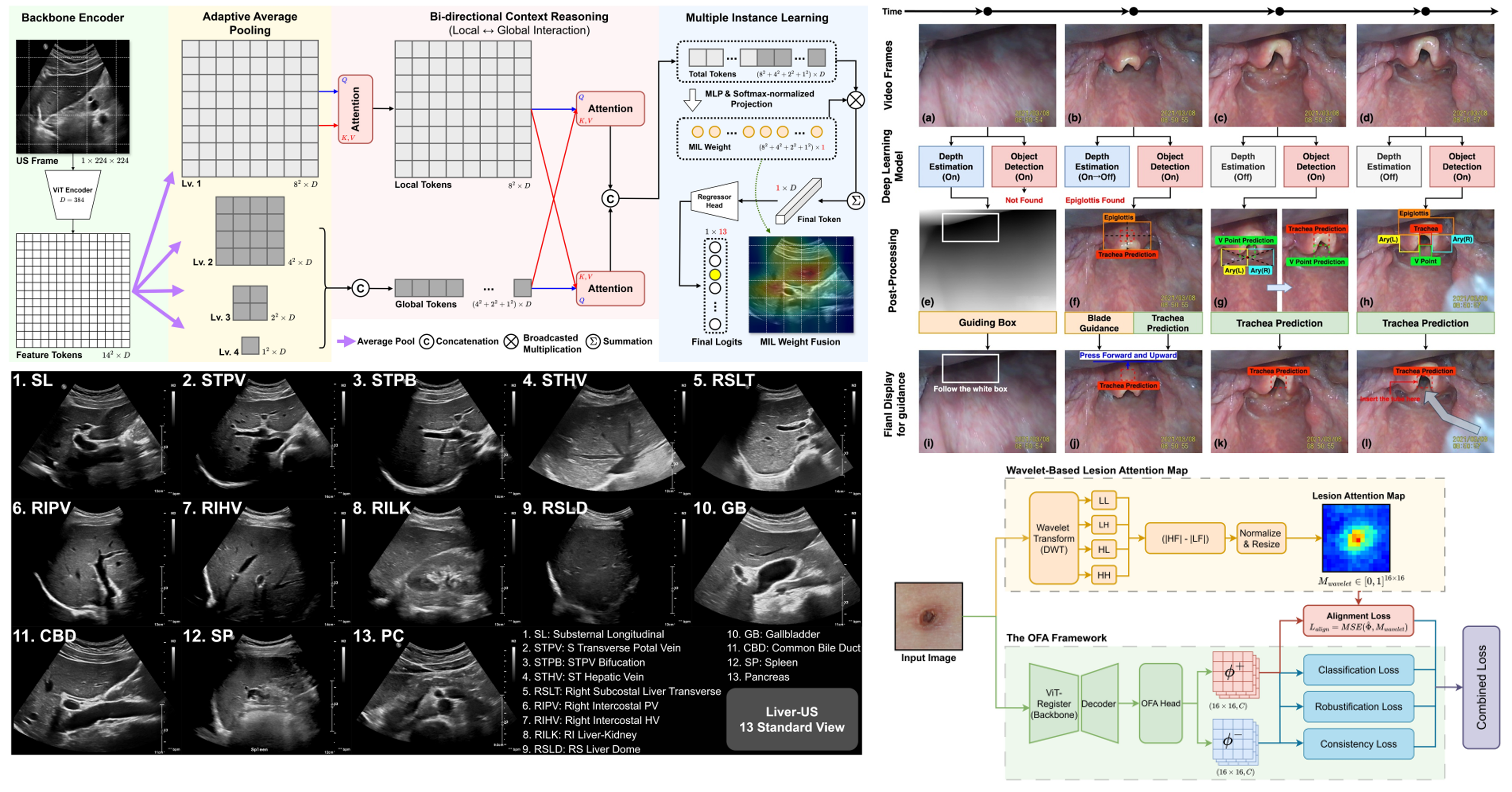
In addition to our core projects, AMILab conducts a broad range of applied research to address diverse problems in real clinical practice. These projects aim to expand the intersection between medical imaging and artificial intelligence, responding to clinical needs such as diagnostic assistance, procedural guidance, disease monitoring, and workflow improvement. Representative projects include abdominal ultrasound view classification, where we develop AI models to identify standard views that are often difficult for novice examiners. This work can directly support ultrasound education and automation, and may also serve as an important source of medical knowledge for future LLM/VLM-based medical AI systems. In our trachea detection project, we are developing an AI platform that automatically detects the trachea and helps guide non-experts during intubation. In our scabies classification project, we develop AI methods to distinguish scabies from other skin diseases using images captured with ordinary cameras, with the goal of enabling more accessible skin disease screening. We are also conducting research on quantitative tracking of disease changes over time. In projects such as longitudinal brain hemorrhage monitoring and longitudinal lung cancer monitoring, we analyze repeatedly acquired medical images to assess changes in lesion size, shape, location, and progression patterns. These technologies aim to support a more precise understanding of disease progression and assist clinical decision-making. These projects are being carried out in collaboration with research teams from various clinical departments at Pusan National University Yangsan Hospital, and represent an important part of AMILab’s translational research efforts centered on real clinical problems.
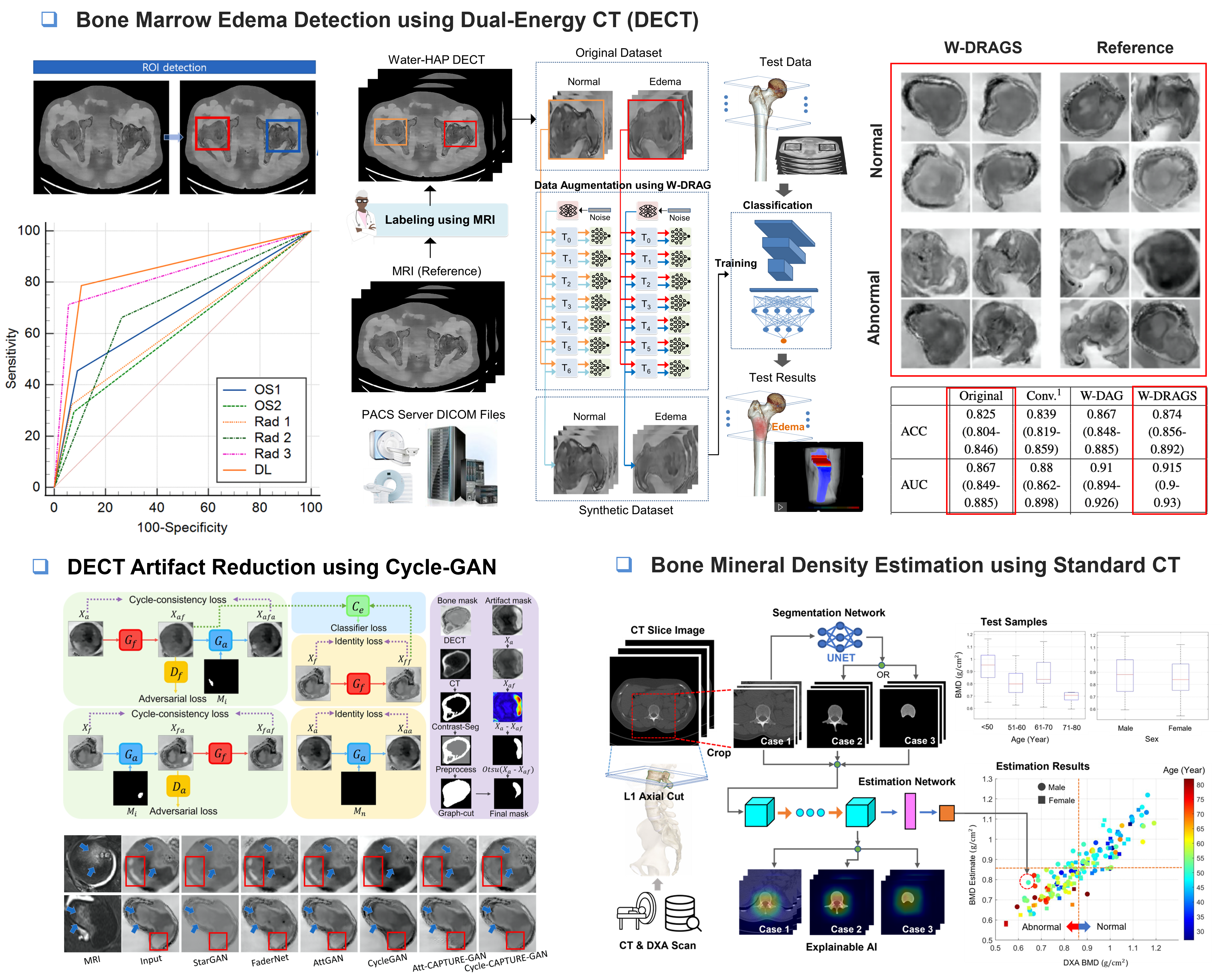
This completed project focused on developing AI technologies for bone marrow edema (BME) detection and bone mineral density (BMD) assessment using musculoskeletal CT images. BME is an important imaging finding associated with trauma, inflammation, fractures, and joint diseases, but it can be difficult to interpret on CT images and may be confused with imaging artifacts. In this project, we developed an AI model that classifies the presence of BME on a slice-by-slice basis using DECT images, enabling more detailed localization of BME within bone rather than simply determining its presence or absence at the patient level. To improve the performance of the BME classification model, we also investigated GAN-based image generation for data augmentation. This approach aimed to enhance model generalization by generating diverse training samples under limited medical imaging data conditions. In addition, to improve radiologists’ BME interpretation, we developed a CycleGAN-based image enhancement model that removes artifacts interfering with diagnosis while preserving true BME-like patterns that may appear similar to artifacts. Furthermore, we developed an AI model that predicts DEXA-based BMD values using conventional CT images alone. This approach suggests the possibility of estimating bone density information from existing clinical CT scans without requiring an additional DEXA examination, potentially supporting more efficient osteoporosis screening and fracture risk assessment. This research was conducted in collaboration with the Department of Radiology at Pusan National University Yangsan Hospital and the Department of Orthopedic Surgery at Busan Medical Center. The project was supported by Pusan National University internal research funding and the AI Convergence Innovation Human Resources Development Program of the Institute of Information & Communications Technology Planning & Evaluation (IITP).